File:Tree of Life nmicrobiol201648-f1 via Nature.jpg
Original file (695 × 833 pixels, file size: 98 KB, MIME type: image/jpeg)
"It's All Connected"
- It's all related, it's all connected
The tree of life is one of the most important organizing principles in biology. Gene surveys suggest the existence of an enormous number of branches, but even an approximation of the full scale of the tree has remained elusive.
Recent depictions of the tree of life have focused either on the nature of deep evolutionary relationships or on the known, well-classified diversity of life with an emphasis on eukaryotes. These approaches overlook the dramatic change in our understanding of life's diversity resulting from genomic sampling of previously unexamined environments. New methods to generate genome sequences illuminate the identity of organisms and their metabolic capacities, placing them in community and ecosystem contexts.
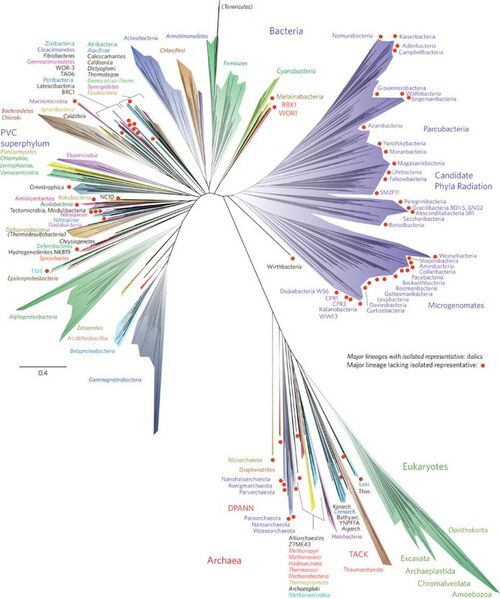
Here, we use new genomic data from over 1,000 uncultivated and little known organisms, together with published sequences, to infer a dramatically expanded version of the tree of life, with Bacteria, Archaea and Eukarya included. The depiction is both a global overview and a snapshot of the diversity within each major lineage. The results reveal the dominance of bacterial diversification and underline the importance of organisms lacking isolated representatives, with substantial evolution concentrated in a major radiation of such organisms. This tree highlights major lineages currently underrepresented in biogeochemical models and identifies radiations that are probably important for future evolutionary analyses.
·····································
"Tiny Little Ones"
Micro-organisms / Microorganisms / Microbiology / Microbiomes
Living Soil, Healthy Soil / Thriving Oceans, Sustainability
• https://www.nature.com/articles/nmicrobiol201648/figures/1
• http://www.nytimes.com/2016/04/12/science/scientists-unveil-new-tree-of-life.html
·····································
The Magic of the Microscopic World
- by Michael Lemonick
• https://blogs.scientificamerican.com/observations/the-magic-of-the-microscopic-world
·····································
Early approaches to describe the tree of life distinguished organisms based on their physical characteristics and metabolic features. Molecular methods dramatically broadened the diversity that could be included in the tree because they circumvented the need for direct observation and experimentation by relying on sequenced genes as markers for lineages. Gene surveys, typically using the small subunit ribosomal RNA (SSU rRNA) gene, provided a remarkable and novel view of the biological world but questions about the structure and extent of diversity remain. Organisms from novel lineages have eluded surveys, because many are invisible to these methods due to sequence divergence relative to the primers commonly used for gene amplification. Furthermore, unusual sequences, including those with unexpected insertions, may be discarded as artefacts.
Whole genome reconstruction was first accomplished in 1995, with a near-exponential increase in the number of draft genomes reported each subsequent year. There are 30,437 genomes from all three domains of life—Bacteria, Archaea and Eukarya — which are currently available in the Joint Genome Institute's Integrated Microbial Genomes database (accessed 24 September 2015). Contributing to this expansion in genome numbers are single cell genomics and metagenomics studies. Metagenomics is a shotgun sequencing-based method in which DNA isolated directly from the environment is sequenced, and the reconstructed genome fragments are assigned to draft genomes. New bioinformatics methods yield complete and near-complete genome sequences, without a reliance on cultivation or reference genomes. These genome-based approaches provide information about metabolic potential and a variety of phylogenetically informative sequences that can be used to classify organisms16. Here, we have constructed a tree of life by making use of genomes from public databases and 1,011 newly reconstructed genomes.
https://www.greenpolicy360.net/mw/images/Tree_of_Life_nmicrobiol201648-f1_via_Nature.jpg
File history
Click on a date/time to view the file as it appeared at that time.
| Date/Time | Thumbnail | Dimensions | User | Comment | |
|---|---|---|---|---|---|
| current | 00:06, 12 April 2016 |  | 695 × 833 (98 KB) | Siterunner (talk | contribs) | http://www.nature.com/articles/nmicrobiol201648 April 2016 http://www.nytimes.com/2016/04/12/science/scientists-unveil-new-tree-of-life.html The tree of life is one of the most important organizing principles in biology1. Gene surveys suggest the e... |
You cannot overwrite this file.
File usage
- Agriculture
- Animals
- Biodiversity
- Biogeosciences
- Biosphere
- Climate Change
- Earth Science
- Earth Law
- Ecological Economics
- Eco-Spirituality
- Ecology Studies
- Endangered Species
- Environmental Full-cost Accounting
- Environmental Laws
- Environmental Protection
- Evolutionary Biology
- Extinction
- Fisheries
- Forests
- Green Graphics
- Green Values
- Land Ethic
- Microbiology
- Microorganism
- Natural Capital
- Natural Resources
- Natural Resource Economics
- Oceans
- Ocean Science
- Ocean Sustainability
- Open Access
- Rights of Nature
- Seventh Generation Sustainability
- Soil
- Sustainability
- Sustainability Policies
- Wetlands
- Zoology